Most explanations of PBIX internals jump straight from “the data is columnar” to “here is the final table.” The missing middle is usually the dictionary layer. VertiPaq does not store every imported value verbatim in the segment payload. It stores encoded IDs and relies on companion structures such as .dictionary and .hidx files to make those IDs meaningful.
That middle layer is where a parser stops being a metadata browser and starts becoming a real decoder.
Where This Fits
This article assumes you already know how to reach DataModel and what the VertiPaq workspace looks like. If not, read The DataModel: Power BI’s Embedded Analysis Services Engine first. If you want the next step after dictionaries, continue with Reconstructing Column Data from .idf and .idfmeta.
Two Main Reconstruction Paths
In practice, imported columns tend to follow one of two broad paths:
- a dictionary-backed path
- a value-encoding path that may also expose a
.hidxfile
The split is visible right in the on-disk filenames. Hash-encoded columns drop a dictionary file into the table folder, typically named #.{Table} ({TableID}).{Column} ({ColumnID}).dictionary. Value-encoded columns drop a hash-index file instead, named #.H${Table} ({TableID})${Column} ({ColumnID}).hidx. That single convention tells you which reconstruction path to take before you open a single byte of the payload.
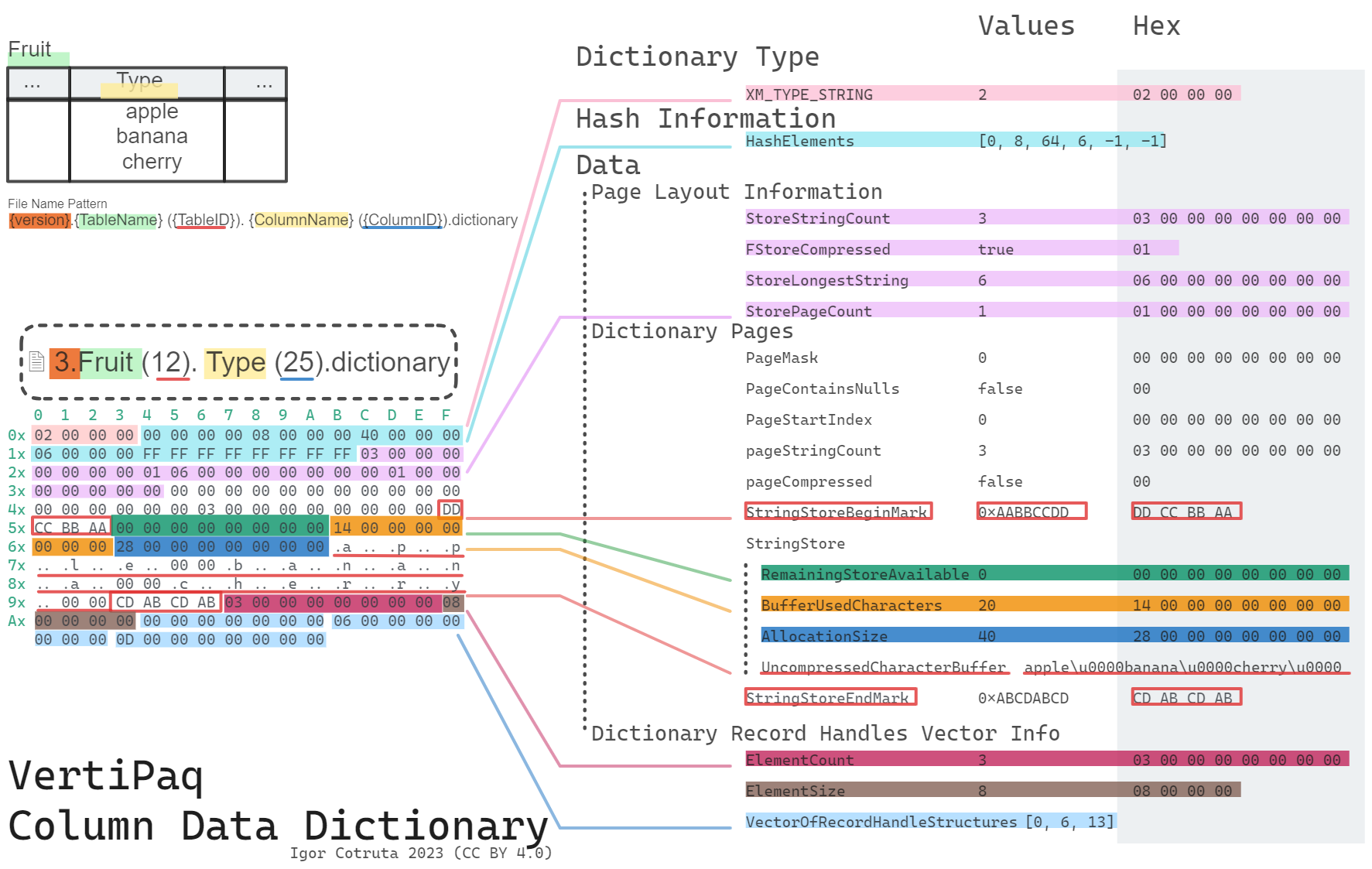
What a .dictionary File Stores
For dictionary-backed columns, the .idf payload stores integer IDs rather than the original strings or numerics. The .dictionary file provides the mapping from internal data IDs back to real values.
The high-level shape is straightforward:
seq:
- id: dictionary_type
type: s4
- id: hash_table_class
type: u4
- id: hash_bin_header
type: hash_bin_headerdictionary_type takes one of four values: -1 (invalid), 0 (int64 values), 1 (float64 values), and 2 (strings). The hash-bin header that follows describes a 64-bin hash table the engine uses in memory — on disk it’s usually empty (with an i64 = -1 sentinel after the three header fields), because the engine rebuilds it from the value store on load. A deserializer that only needs the values themselves can treat that whole block as opaque and skip past it.
Numeric dictionaries are then just a flat vector: an 8-byte count, a 4-byte element size, and then the raw bytes (s4le for 4-byte elements, s8le or f8le for 8-byte elements depending on whether the type is long or real).
String dictionaries are paged. After a small page-layout header (total string count, longest string length, page count) comes a sequence of dictionary pages, each of which can be independently compressed or not based on the low bit of a per-page page_mask. Every page is wrapped by two fixed sentinels — 0xDD 0xCC 0xBB 0xAA at the start and 0xCD 0xAB 0xCD 0xAB at the end — which double as a useful format fingerprint when you’re exploring a file by hand. A final vector of record handles (page_id, bit-or-byte offset) is how the engine locates a specific string within the pages.
Huffman-Compressed String Pages
One of the more interesting findings in the reverse-engineered format is that string pages are not always stored as plain null-terminated text. Compressed pages carry a character_set_type_identifier that selects between two Huffman variants:
0x000aba91— charset-based Huffman. A singlecharacter_set_usedbyte picks a character set (commonly ANSI / Latin-1), and the Huffman tree emits single-byte characters directly.0x000aba92— general Huffman. The Huffman tree emits raw bytes whose sequence is UTF-16LE; there is nocharacter_set_usedfield, and the decoder needs to re-encode its output and decode as UTF-16LE to get real strings.
Each compressed page carries a compact 128-byte encode_array rather than a full 256-byte table. The decoder expands that to 256 codeword lengths (two 4-bit nibbles per byte), reconstructs canonical Huffman codes from the lengths, and walks a tree bit-by-bit to decode. Record handles (page_id, bit-offset pairs) mark where each string starts inside the compressed buffer, so the decoder knows exactly when to stop for each value.
That is why a parser cannot treat the dictionary as a trivial array lookup. It often has to decompress string pages, honor record-handle offsets, and rebuild the mapping page by page. In pbixray, that logic lives in the dictionary-reading path of the VertiPaq decoder.
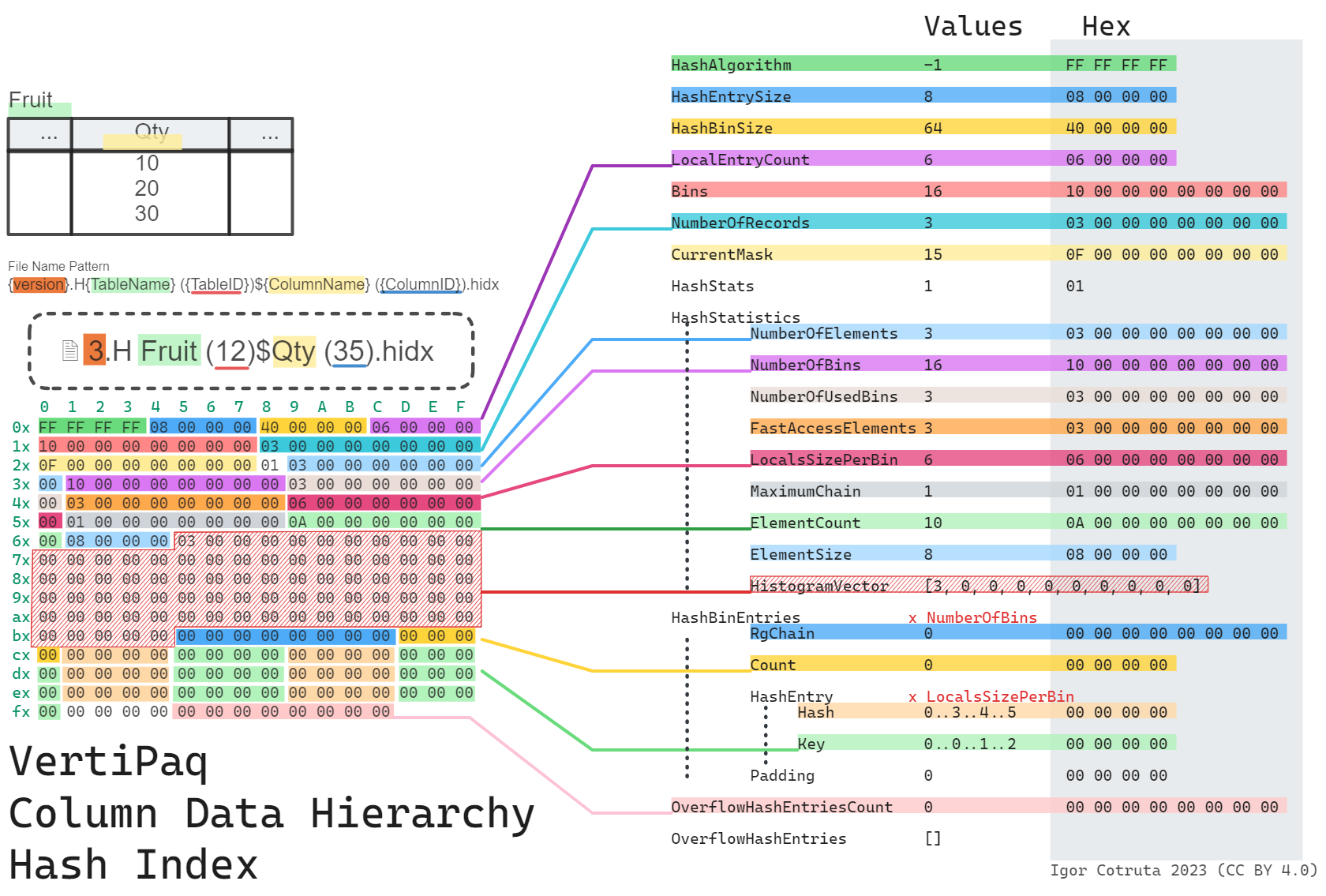
What a .hidx File Stores
A .hidx file is a hash index rather than a value store. Its layout is built around hash bins, local entries, and overflow entries:
seq:
- id: hash_algorithm
type: s4
- id: hash_entry_size
type: u4
- id: hash_bin_size
type: u4
- id: local_entry_count
type: u4The key point is that .hidx supports lookup-oriented behavior around encoded values. It is not the same thing as the dictionary value store.
Internal IDs, min_data_id, and Nullability
The .idf payload is usually decoded into internal IDs first. Those IDs only become user-facing values after one more mapping step.
In the current pbixray implementation, the reconstruction path depends on column metadata:
- if a dictionary file is present, decoded IDs are mapped through the dictionary
- if an
HIDXpath is present instead, values are reconstructed numerically usingBaseIdandMagnitude
That second case is worth underlining. A .hidx file signals a different encoding path, but the actual user-visible number can often be reconstructed from metadata plus the decoded integer vector without consulting the hash index during plain table extraction.
Nullability adds another wrinkle. For dictionary-backed columns, the minimum data ID has to be adjusted correctly so the null slot lands in the expected position.
Why These Structures Matter to a Parser
Without the dictionary and hash-index layer, the output of .idf decoding is only half-finished. You would have:
- integer codes for strings
- encoded offsets rather than final numerics
- no reliable mapping from storage IDs to business values
This is why I think of dictionaries as the hinge between physical storage and semantic output.
How pbixray Uses Them
The current decoder follows a simple pattern:
if pd.notnull(column_metadata["Dictionary"]):
dictionary = self._read_dictionary(dictionary_buffer, min_data_id=meta["min_data_id"])
values = self._read_rle_bit_packed_hybrid(...)
return pd.Series(values).map(dictionary)
elif pd.notnull(column_metadata["HIDX"]):
values = self._read_rle_bit_packed_hybrid(...)
return pd.Series(values).add(column_metadata["BaseId"]) / column_metadata["Magnitude"]That is a useful summary of the whole article. Segment decoding gives you encoded values. The dictionary or value-encoding path gets you back to the values a user would recognize.
Related Articles
- Read Inside
metadata.sqlitedb: Tables, Columns, Measures & Relationships for the metadata that points to these files. - Read Reconstructing Column Data from
.idfand.idfmetafor the segment payloads that produce the internal IDs in the first place. - Read How VertiPaq Sorts Rows to Maximize RLE Compression for the sibling project that explains why row order matters so much for dictionary-backed storage.
- Read Parsing PBIX Files with Python (pbixray) for the practical extraction layer on top of both.